Current projects
Phenomics First – Center of Excellence in Genomic Sciences

As part of a newly established Center of Genomic Science (press release) I am responsible for the unification of phenotype ontologies using community- and pattern-driven ontology development. The ultimate goal is a deeply integrated Unified Phenotype Ontology (uPheno) that enables applications such as cross-species phenotype matching, phenotypic profile matching for disease diagnostics and improved variant prioritisation. I am furthermore responsible for our strategy for ontology merging and mapping, for example by leading the community effort of establishing a Simple Standard for Sharing Ontology Mappings (SSSOM) or working with our stakeholders on the effective utilization of Mouse-Human ontology mappings. Lastly, I am actively contributing to the Mondo Disease Ontology and Human Phenotype Ontology, mainly in the areas of quality control and release pipelines. The Center of Excellence is one of a number of efforts driven by the Monarch Initiative, which seeks to integrate the worlds gene-to-phenotype-to-disease knowledge in a comprehensive knowledge graph to aid variant prioritisation and ultimately clinical diagnostics. Recently, the focus of our efforts started shifting from knowledge graph engineering to graph-based machine learning efforts and large language models. I was first contracted to do this work by EMBL-EBI – a major contributor to open data, open ontologies and open science, and later by Tislab a unique player in Translational research, connecting the clinical and the biomedical research domains.
Augmenting and merging ontologies using ontology mappings and knowledge graph embeddings
It does not matter how hard we try to reconcile our ontologies, for example, as part of efforts such as the OBO Foundry – there will always be some level of overlap between them, i.e. the same terms existing in multiple ontologies. Data is regularly linked to concepts from different (sometimes internal, non-public) ontologies, and this data needs to be integrated – which means that the different ontologies need to be carefully aligned. Furthermore, we want to be able to enrich our own ontologies with information (such as synonyms) from and links to other public sources. Many disciplines play a role in these processes: Ontology and Knowledge Graph Matching is an active research area that seeks to find ways to link terms between ontologies, employing techniques such as string-based matching, graph-based matching or more advanced techniques from Natural Language Processing (NLP). Recently, Machine Learning algorithms and Knowledge Graph Embeddings have been used to greatly enhance traditional mapping techniques, and we expect a lot more to come here. Ontology Merging is concerned with combining two ontologies in a way that the resulting whole is consistent, yet richly axiomatised, and uses approaches from formal logic to Bayesian. Ontology Engineering is the “art” of evolving an ontology in a sustainable way, for example using design patterns, logical reasoning and more classical visualisation approaches focused on manual curation and automated quality control. With SymbiOnt, we try to bring together techniques from all of the above to provide a fully executable workflow for ontology merging and augmentation. The goal is to combine tools and techniques in a Docker-based workflow that enables the end-to-end merging and augmentation of ontologies. For example, we want to be able to combine the many existing disease ontologies and terminologies into a rich coherent framework and keep them synchronised with minimal human curation effort. The work is funded by a gift of the Bosch corporation to the Lawrence Berkeley National Laboratory (LBNL) and involves LBNL and Bosch researchers.
Infrastructure for ontology engineering with Knocean

Toronto-based Knocean is the primary provider of software and services in the area of Open Biological and Biomedical Ontologies (OBO) and a major contributor to our open source tools and infrastructure. Flagship tooling includes for example ROBOT, a powerful system to build ontology pipelines – for years an indispensable item in my knowledge engineering toolbox. Other tools developed by Knocean (or rather the subset I am at least partially involved in) are the OBO-dashboard (a quality control system for all OBO ontologies) and DROID (a UI for executing workflows based on make and git). I am excited to be contracted by Knocean to improve the quality of open ontologies with solid software solutions and being given the opportunity to really make good things better in the wider OBO-sphere.
Past (pre-2021)
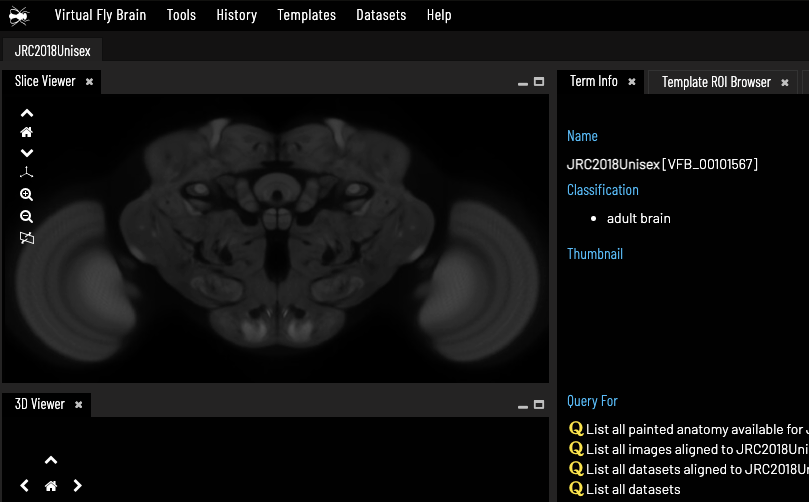
Virtual Fly Brain Knowledge Graph
The Virtual Fly Brain is knowledge graph-driven application for exploring images of neurons, underpinned by a Neo4j knowledge graph. The 3D browser is based on Geppetto, a web-based visualisation platform for building neuroscience software applications developed by Oxford-based MetaCell. The underlying knowledge graph combines a range of OBO ontologies with curated metadata about images generated by Drosophila research around the world, as well as connectomics data. I contributed to the project the design and implementation of a docker based ETL pipeline including processes that convert between OWL and Neo4j in a user-friendly manner and an integration layer based on a triple store behind an rdf4j backend. MetaCell contracted me in 2020 to improve the ETL pipeline and help integrating the new connectomics data.